
In particular, known host receptors include: aminopeptidase N (APN) for 229E angiotensin-converting enzyme 2 (ACE2) for NL63, SARS1, and SARS2 O-acetylated sialic acid ( O-ac Sia) for OC43 and HKU1 and dipeptidyl peptidase-4 (DPP4) for MERS ( 9, 14). Interestingly, the receptor-binding domain (RBD) and host receptor differ among the known human-infecting CoVs. This shows that receptor binding and membrane fusion are important initial and key steps in CoV pathogenesis. In a CoV infection scenario, S1 would bind to a suitable receptor on the host cell surface enabling viral attachment and, subsequently, S2 fuses both the host and viral membranes, thereby, allowing viral genomes to enter host cells ( 9, 10). With regards to the ectodomain, it is further divided into the S1 receptor-binding subunit that mainly functions in viral attachment and S2 membrane-fusion subunit that facilitates virus-cell fusion ( 9, 10). Additionally, the spike protein is comprised of three segments, namely: the large ectodomain, single-pass transmembrane anchor, and short intracellular tail ( 10). Spike protein (a class I viral fusion protein) follows a metastable prefusion conformation upon translation and, likewise, forms trimers that resemble club-shaped spikes along the CoV membrane surface ( 13). This would highlight the significance of the CoV spike protein in terms of viral pathogenesis. Among the structural proteins, the spike protein has been involved in mediating viral entry, determinant of host tropism, inducing viral pathogenesis, and major inducer of host immune responses ( 9– 12). Moreover, CoVs have the largest genome among all known RNA viruses which in-turn is packed in a helical capsid comprised of a nucleocapsid protein (N) and surrounded by a viral envelope which in-turn is associated with structural proteins, namely: membrane, envelope, and spike ( 10). In general, CoVs cause serious health problems to both human and animal hosts and, in particular, CoV infections primarily affect the respiratory and gastrointestinal tracts ( 9). Historically, there are seven known CoVs capable of infecting humans, namely: human CoV (HCoV)-229E (1962), HCoV-OC43 (1967), severe acute respiratory syndrome (SARS)-CoV 1 (SARS1) (2002), HCoV-NL63 (2004), HCoV-HKU1 (2005), Middle East respiratory syndrome (MERS)-CoV (2012), and SARS-CoV 2 (SARS2) (2019) ( 2– 8). Taken together, we propose that the SARS2 spike S1-CTD follows a Pattern III orientation which has a higher degree of similarity with SARS1 and some degree of similarity with both OC43 and HKU1 which coincidentally are in the same phylogenetic cluster and lineage, whereas, the SARS2 spike S1-NTD has some degree of similarity among human-infecting CoVs that are either in the same phylogenetic cluster or lineage.Ĭoronaviruses (CoV) are enveloped positive-stranded RNA viruses that belong to the family Coronaviridae and order Nidovirales with the subfamily Othocoronavirinae composed of four genera, namely: alphacoronavirus, betacoronavirus, gammacoronavirus, and deltacoronavirus ( 1). 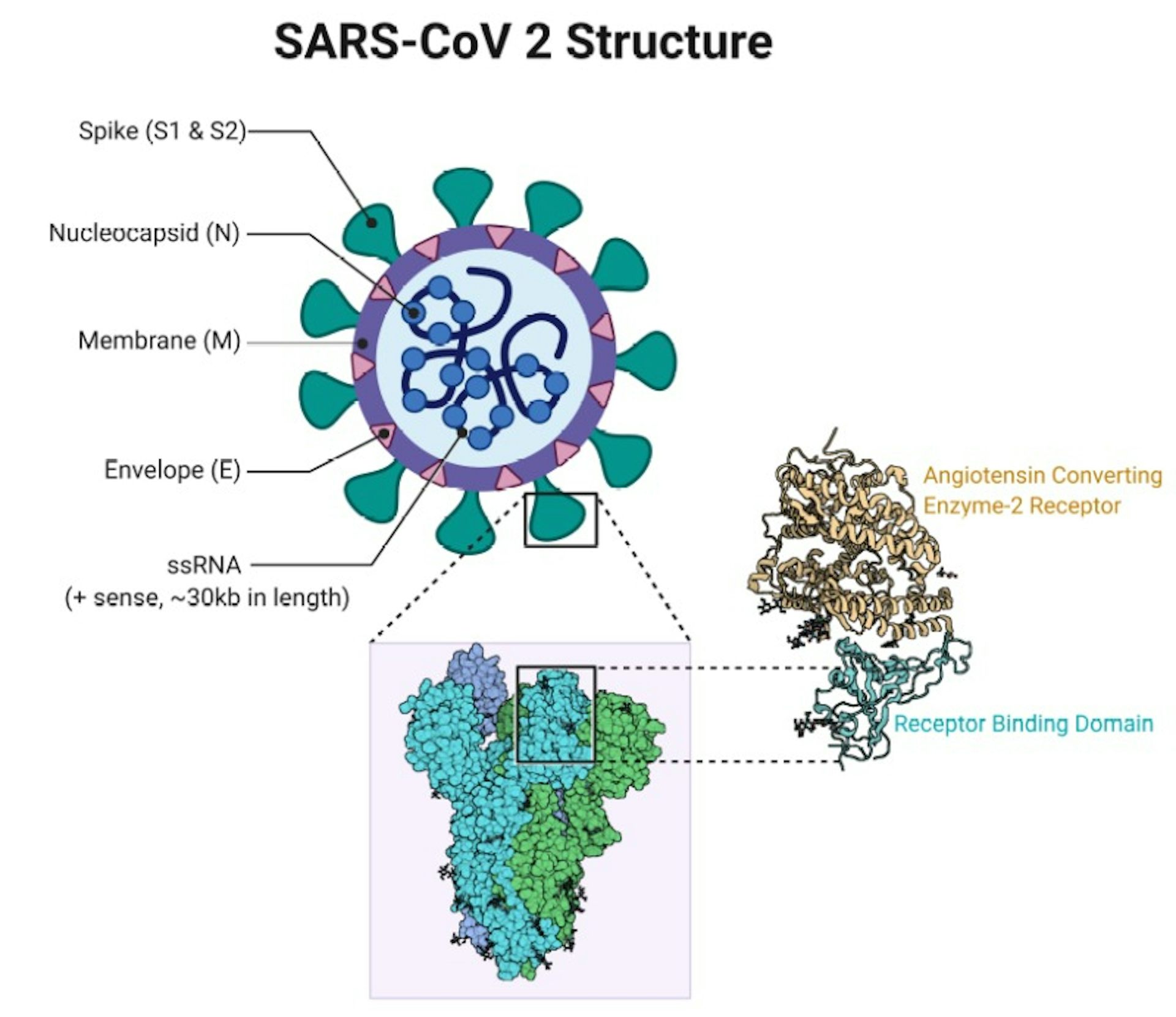
Furthermore, we postulate that human-infecting CoVs in the same phylogenetic cluster may have similar S1-CTD and S1-NTD structural orientation. 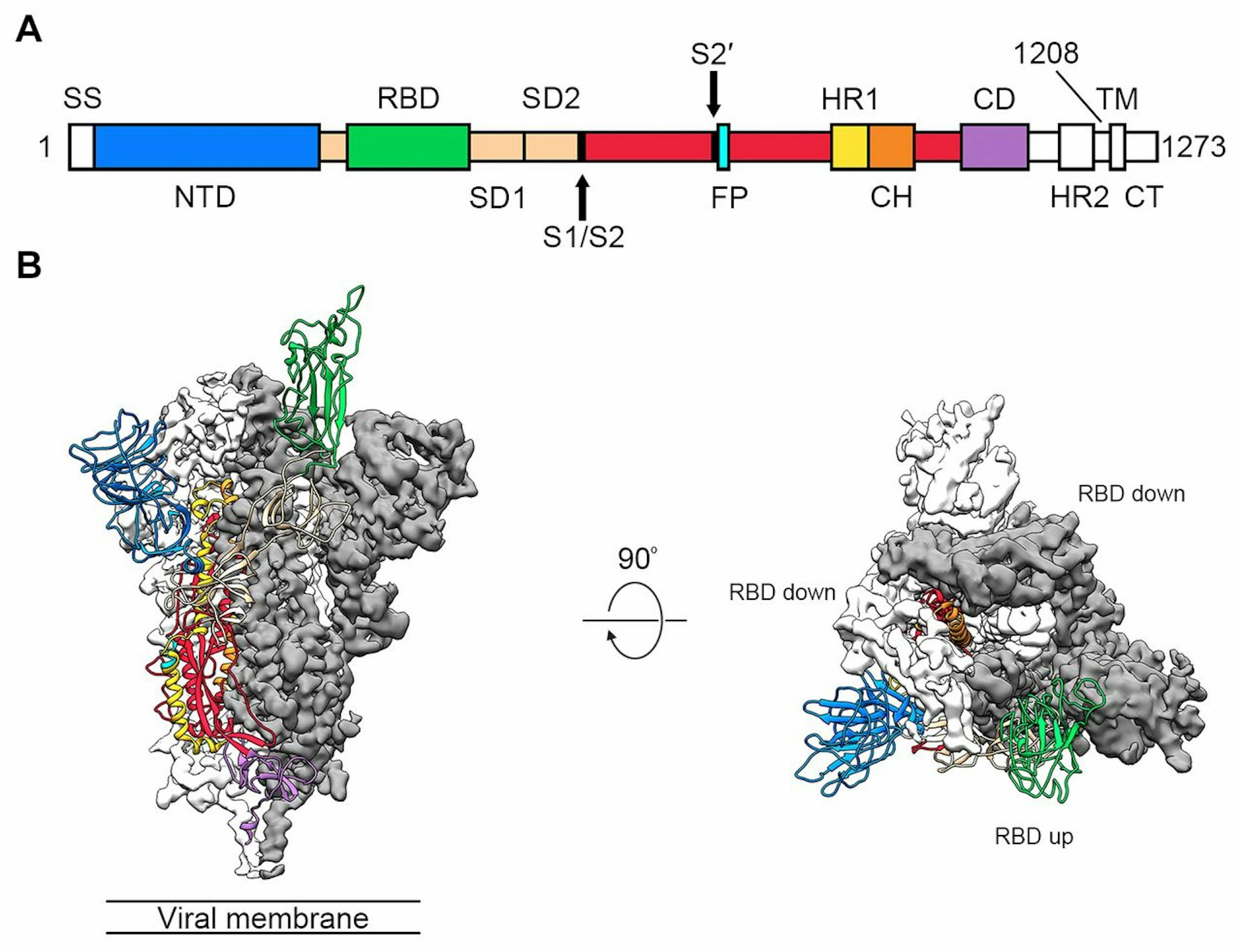
Additionally, we established that there are potentially three distinct S1-CTD orientation (Pattern I, Pattern II, Pattern III) among the human-infecting CoVs. Moreover, we observed that spike models in the same phylogenetic cluster or lineage could potentially have similar protein structure. We found that the structural orientation of S1-CTD is a potential structural feature associated to both the CoV phylogenetic cluster and lineage. Model quality assessment of all CoV models generated, structural superimposition of the whole protein model and selected S1 domains (S1-CTD and S1-NTD), and structural comparison based on RMSD values, Tm scores, and contact mapping were all performed. In this study, we utilized a computational approach to generate models and analyze the seven human-infecting CoV spike proteins, namely: HCoV 229E, HCoV OC43, HCoV NL63, HCoV HKU1, SARS CoV, MERS CoV, and SARS2. Surprisingly, the possible differences and similarities in the protein structural conformation between the various human-infecting CoV spike proteins have not been fully elucidated. Among the human-infecting CoVs, only the infection of SARS CoV 2 (SARS2) among humans resulted to a pandemic which would suggest that the protein structural conformation of SARS2 spike protein is distinct as compared to other human-infecting CoVs. CoV attachment to the host is the first step of viral pathogenesis and mainly relies on the spike glycoprotein located on the viral surface. Department of Microbiology, Nihon University School of Dentistry, Tokyo, JapanĬoronaviruses (CoV) are enveloped positive-stranded RNA viruses and, historically, there are seven known human-infecting CoVs with varying degrees of virulence.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
